We foster dialogue between data scientists and researchers in clinics and laboratories in order to drive excellence in health care research at Stanford.
About the Data Studio
The Data Studio is a collaboration between Spectrum (The Stanford Center for Clinical and Translational Research and Education) and the Department of Biomedical Data Science. The Data Studio is open to the Stanford community engaged in biomedical research. We expect it to have educational value for students and postdocs interested in biomedical data science. The Data Studio features DBDS faculty and staff who offer the following services: workshops, office hours, and one-to-one consultations. When you complete the Data Studio request form, our coordinator and consultants will work with you to choose the right service for your research project. Appointments may be requested by completing the required form.
Workshops are an extensive and in-depth consultation for a Medical School researcher based on research questions, data, statistical models, and other material prepared by the researcher with the aid of our facilitator. During the Data Studio Workshop, the researcher explains the project, goals, and needs. Experts in the related topic from across campus will be invited and contribute to the brainstorming. After the meeting, the facilitator will follow up, helping with immediate action items and summary of the discussion. Ultimately, we strive to pair each PI with a data scientist for long-term collaboration.Office Hours are brief consultations for Medical School researchers during the last session of each month. DBDS faculty are available to advise about your research questions. Consult the schedule below to complete the Office Hour registration form. Once you have registered, you will receive a calendar invitation with the date, time, and location of the session. Bring any data, prior analyses, or other materials that you have. Our consultants may even recommend your project for a Workshop if it is appropriate.
One-to-one consultations for Medical School researchers are available year-round. Our facilitator assigns each request to a data scientist with the relevant expertise.
Partners
General questions about statistical issues may be brought to the STAT390 Consulting Workshop. This is a class offered by the Department of Statistics during each academic quarter that is staffed by graduate students and directed by a faculty instructor. The service typically consists of a single meeting with the researcher to address a specific concern, such as planning of experiments and data analysis. For more information, consult the STAT390 Consulting Workshop web page.
Researchers who are members of the Stanford Cancer Institute (SCI) conducting research projects related to cancer may request assistance from the SCI Biostatistics Shared Resource.
The Genetics Bioinformatics Service Center (GBSC) offers an end-to-end bioinformatics consulting service (BaaS) that provides high performance computational infrastructure and cutting-edge bioinformatics services for the Stanford community. The team consults on genomics, transcriptomics, proteomics, epigenetics, and metabolomics projects, and also develop custom workflows. For consulting and hands-on bioinformatics help with your projects please reach out to gbsc-baas-team@lists.stanford.edu to set up an initial meeting.
Schedule
The Data Studio is held each Wednesday from 3:00 until 4:30 pm during the fall, winter, and spring quarters of the academic year. Consult the schedule below for the location of each session. Students may participate by enrolling in BMDS 291 for an introduction to the art of statistical consultation and practicum working on projects with a biomedical researcher. All are welcome to attend. Click here to sign up for our mailing list.
The currently scheduled topic is listed below.
TITLE: Mismatched End (MEnd) Ligase DNA Complex
INVESTIGATORS:
Chun Tsai (1)
Andrey Malkovskiy (2)
Gilbert Chu (3)
(1) Howard Hughes Medical Institute
(2) Carnegie Institute for Science, Department of Plant Biology
(3) Division of Oncology, Department of Medicine
DATE: Wednesday, 8 April 2026
TIME: 3:00–4:30 PM
LOCATION: Room R358, Edwards Building, 300 Pasteur Drive, Stanford, CA
WEBPAGE: https://dbds.stanford.edu/data-studio/
ABSTRACT
The Data Studio Workshop brings together a biomedical investigator with a group of experts for an in-depth session to solicit advice about statistical and study design issues that arise while planning or conducting a research project. This week, the investigator(s) will discuss the following project with the group.
INTRODUCTION
The research is a study of non-homologous end-joining repair mechanism of DNA strands. Nonhomologous end joining (NHEJ) is the primary pathway for repairing DNA double-strand breaks (DSBs) in mammalian cells. DSBs can be created by exogenous agents such as ionizing radiation or by the endogenous mechanism for generating immunological diversity, V(D)J recombination. NHEJ senses broken ends, aligns and joins the ends while optimizing the preservation of DNA sequence. The expectation is that a DNA first binds a Ku-protein that is anchored to the end of a pure DNA strand (image group A). Following that, XL and XLF enzymes (MEnd ligase complexes) bind to the Ku protein in alternating fashion, forming a functionalized DNA tail (image group B) that can now be aligned to a tail of a different DNA strand. This alignment facilitates binding of two DNA strands at their ends.
HYPOTHESIS & AIM
DNA damage caused by internal metabolic processes (oxidative stress, errors during DNA replication) and external environmental factors (toxic chemicals and ionizing radiation, viral load) leads to structural alterations in the molecule. This can cause single- and double-strand DNA breaks. The latter can be repaired in ongoing cell maintenance via the Non-Homologous End Joining (NHEJ). A critical component of this repair is MEnd ligase complex. The purpose of the study is to determine the spatial arrangement of these complexes for the start of the repair process using atomic force microscopy (AFM) topographical images of DNA with Ku protein attached, as well as complete DNA-Ku complexes with XL and XLF ligases. AFM imaging allowed us to take a snapshot of the binding reaction with structural details and to support other indirect experimental findings. We have several hypotheses that AFM was used to address:
- Ku binding to DNA is distributive, but a substantial fraction of Ku molecules are bound to the ends
- The DNA-Ku intermediate and complete complexes prove the formation of filamentary complexes rather than bundles
- Height and width data at the location of Ku binding sites can show if there is preferential orientation of bound Ku
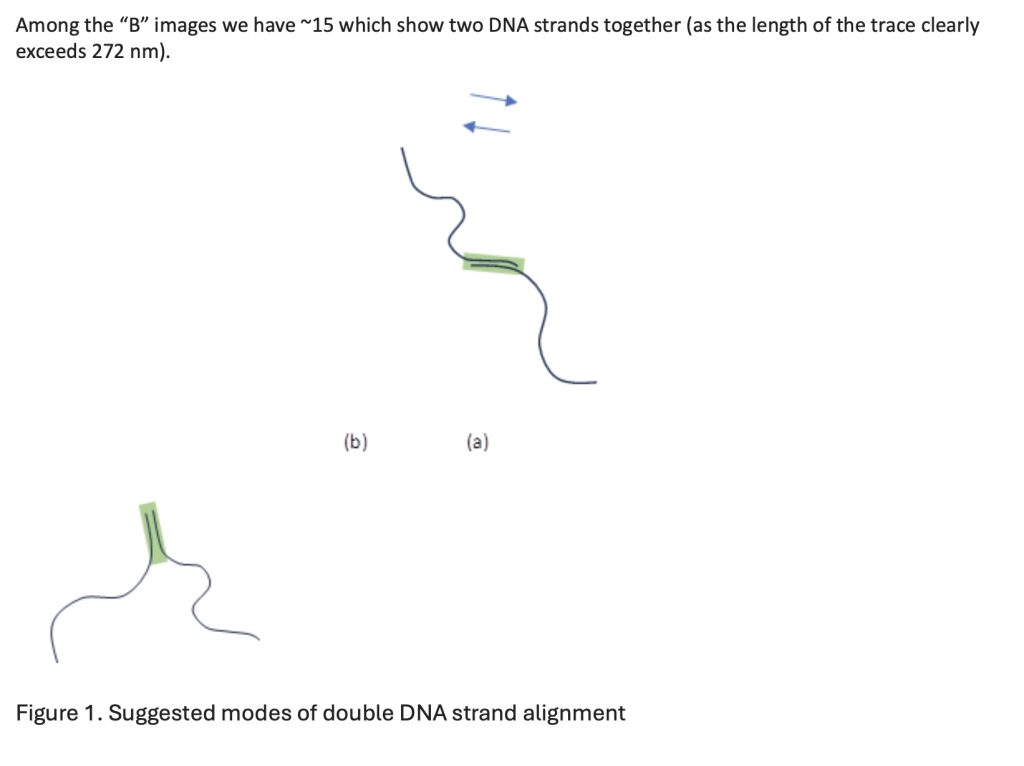
- Ku:XL:XLF complex filament wraps around double-stranded DNA, extends to overhangs and facilitates direct joining. In this case, the ends of DNA are aligned in the same direction, parallel to each other (Figure 1a).
DATASET
We have multiple ~25 images of DNA-Ku intermediate (group A) and 81 images of DNA-Ku full complex (group B). Each 2D topographic (AFM) image represents a DNA complex spread on the surface of a Si wafer. The immobilization is done via covalent 3-Glycidyloxypropyltrimethoxysilane (GPTMS) binding of C-termini of Ku protein and/or XL and XLF enzymes to the Si wafer. No pure DNA strands are bound and are washed away during sample preparation. For “A” the surface is only covered with a layer of GPTMS, which has RMS roughness of ~0.2 nm and is generally very uniform (“mushroom” regime). For “B” the entire surface is covered with XL and XLF complexes, which are rougher in topography. On top of that sit the complexes.
To measure the height and width of DNA-complex traces we have developed our own software, based on a similar one in Ref.1. The inherent error for height estimates is much lower than that for widths due to the nature of the AFM technique. The reason for the development was multi-fold. First, the estimation of widths as performed by code in Ref.1 is not correct due to errors in the code. Second, we have implemented over-sampling of the data, which results in higher resolution of the traces. Finally, we were able to test two different approaches to DNA folding on the surface: stick segments and continuous manual tracing. The latter results in 1.223 times longer traces and agreed better with theoretical predictions of free length of the DNA segments used in the study (300 nm).
STATISTICAL MODELS
We would like to see if there can be other statistical methods of a simple density-based calculation (average height of DNA complex next to the expected point of end-joining).
STATISTICAL QUESTIONS
(1) What is the best method to assess the degree of co-alignment of DNA molecules for the repair process?
(2) What is the best way to calculate the length of the green region (the area of XL-XLF complex sitting on the DNA)? We assume this should be done with lone (non-joined) DNA complexes, of which we have ~65 traces.
(3) How powerful can be this analysis given the limited number of observations of each conformation?
ZOOM MEETING INFORMATION
Join from PC, Mac, Linux, iOS or Android: https://stanford.zoom.us/j/92414292941?pwd=9sQzfFbJpC71PyS5kRofKsT86nEWD9.1
Password: 124320
Or iPhone one-tap (US Toll): +18333021536,,92414292941# or +16507249799,,92414292941#
Or Telephone:
Dial: +1 650 724 9799 (US, Canada, Caribbean Toll) or +1 833 302 1536 (US, Canada, Caribbean Toll Free)
Meeting ID: 924 1429 2941
Password: 124320
International numbers available: https://stanford.zoom.us/u/aMbftTO9
Meeting ID: 924 1429 2941
Password: 124320
Password: 124320